Estimated read time: 3-4 minutes
This archived news story is available only for your personal, non-commercial use. Information in the story may be outdated or superseded by additional information. Reading or replaying the story in its archived form does not constitute a republication of the story.
SALT LAKE CITY — Imagine recording everything you do in your life. Then imagine digging through tens of thousands of entries of data, trying to make sense of it all.
With the recent advancement of genetic sequencing technology, scientists have experienced an explosion of information that, if analyzed, could lead to innovations in biology, find cures for diseases, and so much more. The challenge is finding a way to sort through massive amounts of data, trying to find patterns and connections.
Some scientists call it "drowning in data."
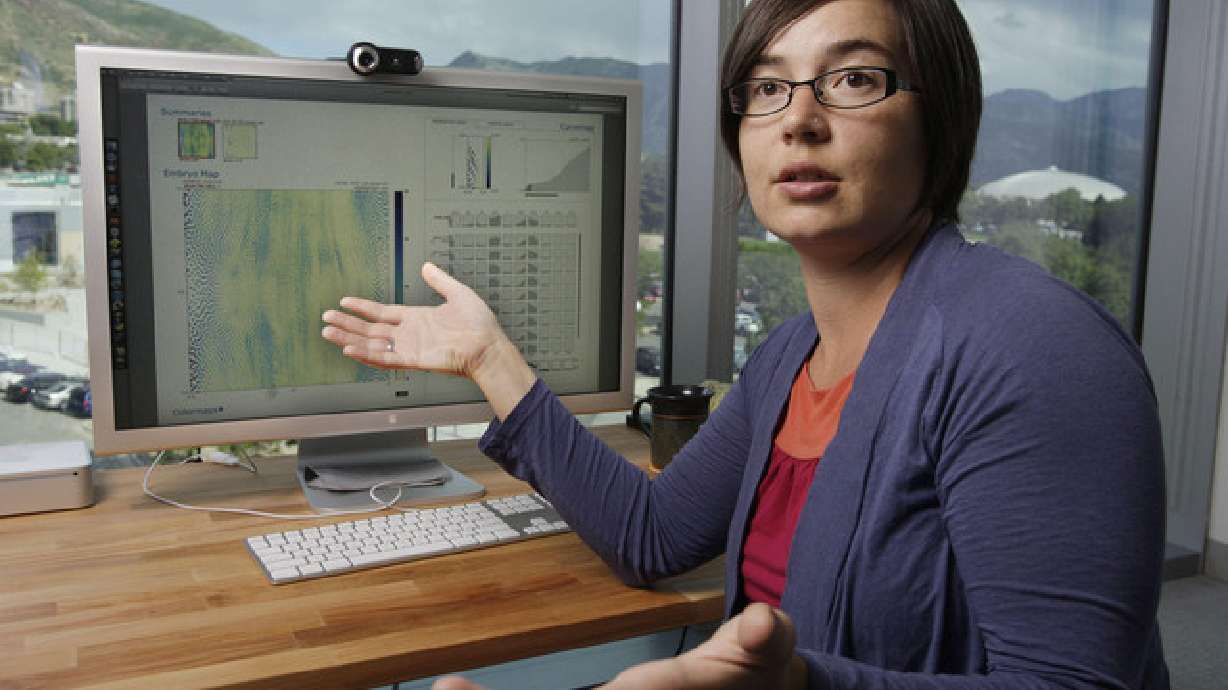
Miriah Meyer recalls talking to biology researchers while doing post-doctorate work at Harvard University and seeing how undeveloped the data software they were using was. "It was shocking to me how much room for improvement there was," Meyer said.
Now a USTAR assistant professor of computer science at the University of Utah, Meyer has worked to create a suite of software tools that helps scientists visualize their data in ways that will help them discover patterns and genetic connections.
It was shocking to me how much room for improvement there was.
–Miriah Meyer
Because of her work, Meyer was recently recognized by the MIT Technology Review's top 35 innovators of 2011. The list honors some of the most innovative researchers in the country.
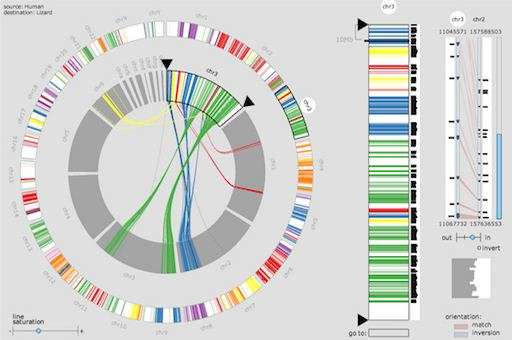
Meyer points to a wheel of colors. What looks like a rainbow of tiny colored threads sprout from the center to the edge of the wheel. Called MizBee, the software helps scientists compare the genomes of one species, in this case the chromosome of a stickleback fish, to another species, like a blow fish. Each color on the wheel represents a gene and the tiny threads indicate a genetic link between species. By translating data into a simple visual image, researchers can easily trace the genetic sources of animal traits.
Angela DePace, a Harvard biologist who has worked with Meyer, said creating software that is made specifically for them will open up more possibilities for discovery. "More often than not, biologists make do with out-of-the- box solutions that are difficult to tailor to their needs," DePace said.
Another tool, called MulteeSum, shows a wave of multicolored dots. Each dot represents the cell of a fruit fly embryo. Using this tool, a scientist can compare the genome of 12 different species of fruit fly. Although the genes are the same, each fly's genetic switches are timed differently, resulting in different traits, Meyer said.
The color difference in the software helps tell the scientist when the activation of certain genes in cells are similar, or different. By clicking on a single cell, a chart of gene activation among the 12 fly species is pulled up.
Her appointment to the University of Utah was made possible through funding from the Utah Science Technology and Research initiative, a long-term, state-funded program that recruits world-class researchers to Utah's universities and funds state-of-the-art research.
Meyer said she hopes her work will help biologists better harness large amounts of data. She said the tools have been released as open-source software, available to scientists all over the world for free.
"We encode meaning through images," Meyer said. "That offers a ton of power."
Email: gfattah@desnews.com