Estimated read time: 1-2 minutes
This archived news story is available only for your personal, non-commercial use. Information in the story may be outdated or superseded by additional information. Reading or replaying the story in its archived form does not constitute a republication of the story.
Tonya Papanikolas ReportingScientists from around the world have been trying to map the most common genetic differences in human DNA. Today those scientists convened in Salt Lake to announce the map's completion.
If you took two random people, their DNA would be 99.9 percent identical. But that 0.1 percent difference is what makes them unique and is very important to researchers.
Hundreds of researchers have been working on this mapping project for three years. They collected blood samples from 269 volunteers in four different regions of the world, including Africa, Japan, China and right here in Utah. DNA from those people was then analyzed to find differences.
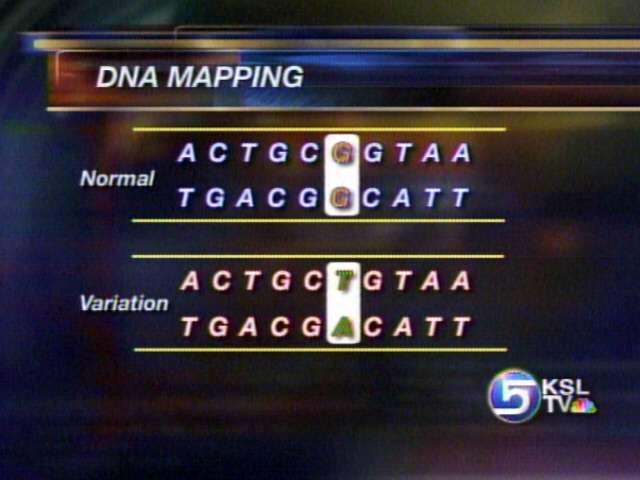
The majority of DNA follows a certain chemical sequence, but sometimes one of those chemicals will variate. Researchers were able to take a look at the variations from the samples collected. These variations, called SNPs, may be able to explain why a person is more susceptible to a certain disease or why some people respond differently to drugs.
Already scientists have used this data to uncover a variation that increases the risk of macular degeneration in the elderly.
Mike Leavitt, U.S. Health & Human Services Secretary: “This is an exciting possibility. We now move toward possible preventions that could save sight and save life. It is a profound step forward, a triumph for collaborative science."
Mark Leppert, Human Genetics Professor, U of U: “The writing of prescriptions in the future will hopefully be one in which personal genetic variants will be tested for and the physician will have a much more sophisticated way of prescribing medications."
Researchers found more than a million different SNPs in phase one of their mapping project. Phase two is adding an additional two-million snips to analyze. All of these are being transferred to public databases.