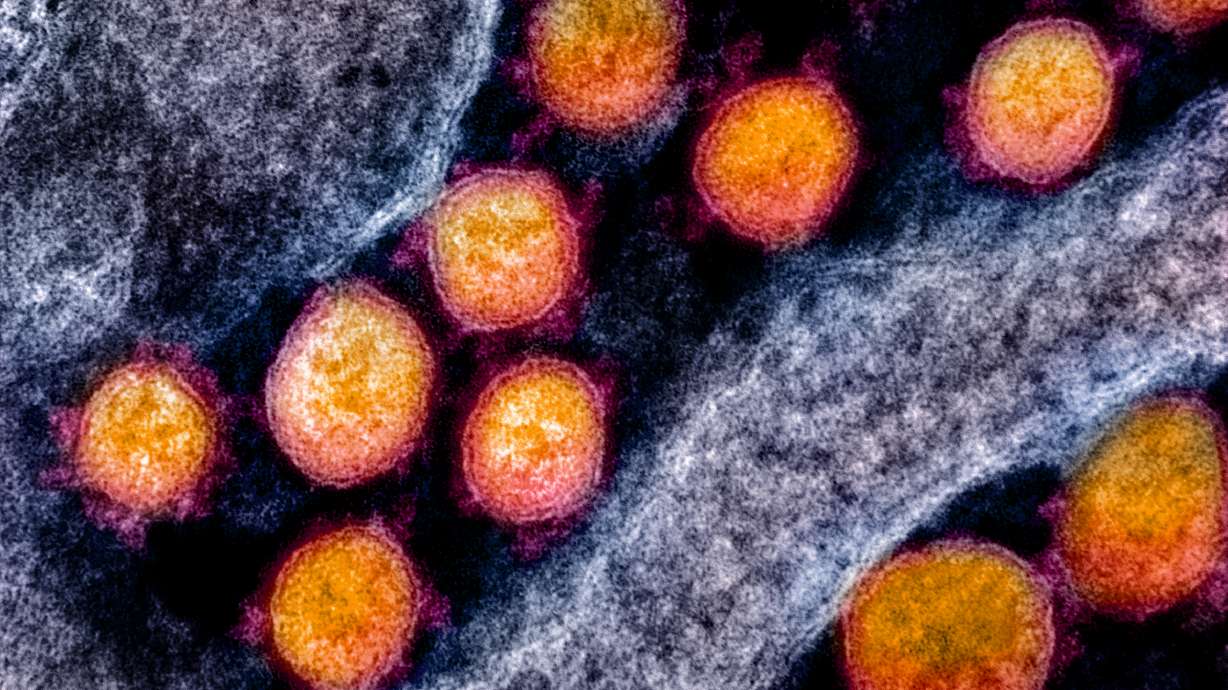Estimated read time: 5-6 minutes
This archived news story is available only for your personal, non-commercial use. Information in the story may be outdated or superseded by additional information. Reading or replaying the story in its archived form does not constitute a republication of the story.
SALT LAKE CITY — In the art of war against plagues and pathogens, the proverb about knowing your enemy has submicroscopic relevance.
As communities unite against the COVID-19 pandemic by keeping a safe distance, researchers have been busy in their labs getting to know everything they can about the SARS-CoV-2 virus causing it.
The history of coronaviruses
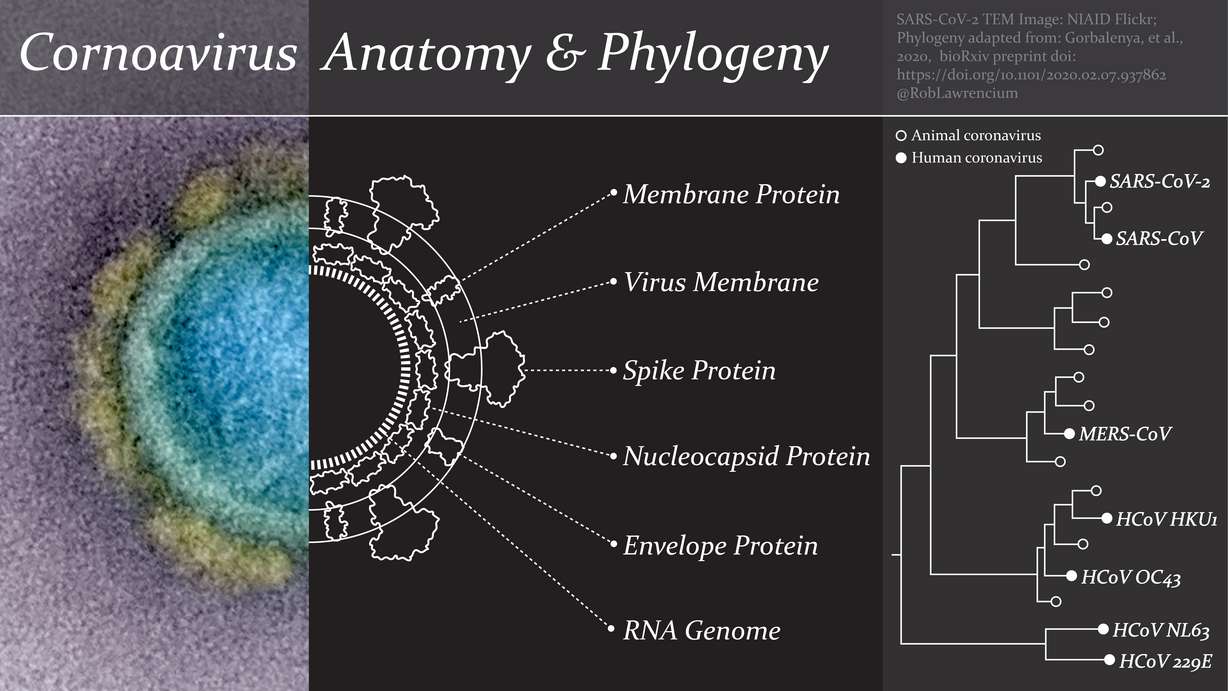
SARS-CoV-2 is the newest member of the coronavirus family. Information about this group of viruses has its roots in the mid-1960s when the first two human coronaviruses were isolated from patients with symptoms of the common cold. Those viruses were named OC43 and 229E.
Other coronaviruses were identified in different mammals and birds during the following decades, and these caused minimal concern until one of them made the jump to humans in 2002. This was the SARS coronavirus, and it took the lives of 774 people before it was quarantined into extinction by 2004.
In the wake of SARS, there was a surge of research interest in coronaviruses that led to the discovery of two more cold-causing human coronaviruses named HKU1 and NL63.
Attention to coronaviruses then waned until 2012, when the MERS coronavirus made the jump from camels to humans causing major outbreaks in Saudi Arabia and South Korea. MERS persists at low levels today and has been the cause of at least 858 deaths so far.
The emergence of SARS-CoV-2 this year brings the count of human coronaviruses up to seven, in addition to dozens of animal coronaviruses.
The characteristics of coronaviruses
These coronaviruses all share some common characteristics. Their shape is round but not necessarily spherical, and they can measure somewhere between 100 to 200 nanometers in diameter. For perspective, if you could line up a row of coronaviruses next to each other, at least a couple thousand could fit between the ridges of your fingerprint.
Like all viruses, coronaviruses are a genome wrapped inside a set of duplicate protein units that have complex three-dimensional shapes and biological functions. And like some viruses, coronaviruses are also encapsulated by a greasy lipid membrane.
Microscope images of coronaviruses show a ring of globs extending from the perimeter, giving the appearance of a crown (or "corona"). These globs are called spike proteins, and they help the virus cling to receptors on specific cell types to initiate infection. Mutations that lead to tiny changes in the composition of these spike proteins can enable a coronavirus to infect a new host, including humans.
The spike protein triggers an immune response when it binds to a cell, so knowing details about the spike composition is useful for designing a vaccine that can trigger the same kind of response. Structural biologists have already produced detailed molecular models of some SARS-CoV-2 proteins including the spike, and now at least 15 SARS-CoV-2 vaccines are in the development pipeline.

Killing SARS-CoV-2
The coronavirus spike proteins are rooted in a membrane of lipids that surrounds the virus. That membrane also holds other important structural units called envelope proteins and membrane proteins, which have roles in the assembly and stability of the virus.
In the same way that oily and greasy residues on dirty dishes dissolve with soapy water and friction, the fatty virus membrane also falls apart when washed with soap and water. The alcohol in hand sanitizer has the same effect. And when the membrane falls apart, so does the virus.
Beneath the membrane and at the core of a coronavirus is its genome, which happens to be quite large and prone to mutation. Coronavirus genes are coded by a long strand of RNA that is wrapped around nucleocapsid proteins for stability — sort of like a string of 30,000 Christmas lights tangled into a ball among tree ornaments.
Lab tests currently being used for SARS-CoV-2 are designed to detect a specific segment of this RNA in nasal swabs taken from patients.
Once the coronavirus infects a cell and hijacks its biology, its genes will encode instructions for making new copies of the RNA genome and protein units. Those units assemble inside the infected cell’s membranes forming new viruses that are released and continue the chain of infection.
The genome also encodes roughly 20 other proteins that orchestrate this process but aren’t part of the virus’s anatomy. Some of these non-structural coronavirus proteins are the targets of antiviral therapeutics currently in development. Ideally, such drugs would bind specifically to those proteins and block their function.
This foundation of knowledge we have about coronaviruses is the result of detailed scientific work that has trickled, ebbed or flowed out of research labs in the last 55 years. So far this year, scientists and physicians have added a flood of new findings about SARS-CoV-2 to that knowledge base at an unprecedented rate.
What they conclude from those findings will be vital, not only for halting this pandemic, but preventing the next one.